PNPLA3 and nonalcoholic fatty liver disease: towards personalized medicine for fatty liver
Nonalcoholic fatty liver disease (NAFLD) affects approximately 25% of the global population and the future clinical and economic consequences are enormous. The rising disease burden from NAFLD reflects increased rates of obesity and its metabolic consequences, but genetic factors clearly determine how individuals respond to excess caloric intake and the resulting metabolic derangements. The review by Barbara and colleagues (1) focuses on key genetic variants of NAFLD from an ever-growing list (2). Of all the genetic variants, patatin-like phospholipase domain-containing protein 3 (PNPLA-3) has emerged as a major common determinant of NAFLD and may be envisioned as part of a future personalized management strategy.
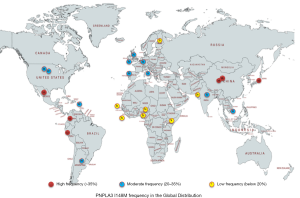
The PNPLA3 story begins in 2008, when a genome-wide association study of the Dallas Heart Study cohort identified a significant association between rs738409 C>G, encoding Ile148Met (I148M) and NAFLD, independent of BMI, diabetes, and alcohol use (3). In this ethnically diverse North American cohort, the risk allele (*I148M) was most common in Hispanics, intermediate in those of European descent and lowest in African Americans. Fast forward to 2019, with numerous GWAS and candidate-gene studies having firmly established PNPLA3 I148M as a genetic modifier of hepatic steatosis and a risk factor for steatohepatitis, fibrosis, and hepatocellular carcinoma (HCC) (4). The distribution of risk allele varies globally, with those of Hispanic and Asian ancestry more likely to carry the M variant (5) (Figure 1).
The magnitude of genetic influence of PNPLA-3 on liver-related outcomes is noteworthy. A meta-analysis with the total of 14,266 NAFLD subjects, the M-variants (GG and GC) were associated with a 3.24-fold and 2.14-fold higher odds of HCC than those with those homozygous for wild-type (CC) (6). In a well-characterized biopsy-proven NAFLD cohort from northern European with matched NAFLD-HCC cases to non-HCC NAFLD cases, the M-variants conferred a 2.26 higher risk of HCC risk than those with wild-type (7). Increasing body mass index amplifies the risk of the 148M-allele, with a study from Copenhagen showing the odds of cirrhosis among persons with BMI >35 kg/m2 was 5.8 in homozygotes for the M-variant (versus wild-type) and corresponding odds of cirrhosis for those with BMI <25 kg/m2 was 2.4 (8). In a large Italian cohort, PNPLA3 M-variants were independently associated with a higher risk of decompensation [hazard ratio (HR), 2.04], HCC (HR, 2.66), and liver-related death (HR, 3.64) among biopsy-proven NAFLD patients (9). Thus, epidemiologic data support PNPLA3 I148M as a major genetic modifier of NAFLD.
PNPLA3 is a crucial lipid regulator in hepatocytes and stellate cells. PNAPL3 is regulated by TGF-β to release retinol from retinyl esters in stellate cells, involving stellate cell activation and fibrogenesis. Within hepatocytes, PNPLA3 is located in the lipid droplets, hydrolyzing triglycerides (TG) and catalyzing the transfer of the polyunsaturated fatty acids (PUFA) from di- and tri-acylglycerols to phosphocholines (10). Its function is crucial in remodeling phospholipids of lipid droplets. Degradation of PNPLA3 is through ubiquitylation of lysine and targeted to the proteasomal degradation. The PNPLA I148M mutant is impaired in hydrolysis activity and less accessible for ubiquitylating degradation, which lead to retention of triglycerides and PUFA enriched lipid droplets, priming accumulation of liver fat. NAFLD patients with the 148M/M variant have higher levels of liver fat, lower serum fasting triglyceride levels, and are less insulin resistant. Interestingly, the retention of PUFA TGs in the liver and decease in the efflux of PUFA into VLDL-TG particles may mitigate cardiovascular disease in PNPLA3 148M carriers. Thus, a genetic profile favoring beneficial outcomes for one aspect of the metabolic syndrome may be detrimental for another.
Since accumulation of I148M mutants alters lipid composition in hepatocytes and stellate cells and “primes the pump” in terms of NAFLD and its sequelae, a therapeutic approach that directly targets 148M is appealing (11). In a homozygous PNPLA3 148 M/M mutant knock-in mice model with the human PNPLA3 I148M mutation introduced into the mouse PNPLA3 gene using homologous recombination, silencing PNPLA3 with a GalNAc3-conjugated-antisense oligonucleotides (ASO) markedly improved NAFLD activity score (NAS) and fibrosis stage in PNPLA3 mutant knock-in mice (12). PNPLA3-ASO also exerted some beneficial effects in wild type mice, likely due to nonselective suppression of wild type PNPLA3. These preclinical studies suggest new therapeutic targets, focused on reducing the mutant PNPLA3 protein, and where screening for PNPLA3 polymorphisms would be relevant for a personalized medicine approach.
Of more immediate clinical relevance is the association PNAPLA3 genotype with response to lifestyle interventions. Patients with I148M mutants are more responsive to carbohydrate and calories restriction, while less responsive to omega-3 fatty acid and statin treatment. In a post-hoc analysis of 154 NAFLD patients from Hong Kong randomized to low calorie diet versus routine care for 12 months, I148M carriers in the dietary intervention group showed greater reduction of hepatic TG, body weight, central obesity and total cholesterol than those without I148M (13). Similarly, in a Polish randomized trial of stringent calorie restriction (500-Kal diet), the homozygous 148M/M had a greater reduction of hepatic steatosis compared to 148I/I (14). In a randomized controlled trial of high dose omega-3 fatty acids for 15–18 months in adults with NAFLD, PNPLA3 148 M/M genotype was independently associated increased liver fat; 10% higher at study end than those with either PNPLA3 148I/I or 148I/M (15). Finally, in a large European cohort with biopsy-proven NAFLD, statin use was associated with lower rates of steatosis, steatohepatitis and fibrosis overall, but PNPLA3 genotype was an important modifier, with benefits only seen in those without the 148I/M variant (16). Thus, NAFLD treatment interventions, whether lifestyle or medications, may be influenced by PNPLA3 genotype. This means that stratification for PNPLA3 I148M variant status in all therapeutic trials is essential to avoid negative results due to this confounder.
The review by Barbara and colleagues highlights the complexity of NAFLD and the strong interplay between genetics and environment (1). The drug pipeline for new NAFLD therapy is rich and targets metabolic, inflammatory/oxidative stress and fibrogenesis pathways. Additional novel therapies related to an understanding of genetic modifiers such as PNPLA3 are important to consider. Ultimately, incorporation of NAFLD genetic modifiers into future clinical risk prediction tools and treatment algorithms may be anticipated enhance patient outcomes.
Acknowledgments
Funding: None.
Footnote
Conflicts of Interest: Both authors have completed the ICMJE uniform disclosure form (available at https://hbsn.amegroups.org/article/view/10.21037/hbsn.2019.10.35/coif). NAT, MD reports grants from Gilead Sciences, personal fees from Intercept, personal fees from EXIFI Management Consultants, LLC, outside the submitted work. The other author has no other conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Barbara M, Scott A, Alkhouri N. New insights into genetic predisposition and novel therapeutic targets for nonalcoholic fatty liver disease. Hepatobiliary Surg Nutr 2018;7:372-81. [Crossref] [PubMed]
- Eslam M, Valenti L, Romeo S. Genetics and epigenetics of NAFLD and NASH: Clinical impact. J Hepatol 2018;68:268-79. [Crossref] [PubMed]
- Romeo S, Kozlitina J, Xing C, et al. Genetic variation in PNPLA3 confers susceptibility to nonalcoholic fatty liver disease. Nat Genet 2008;40:1461-5. [Crossref] [PubMed]
- Anstee QM, Day CP. The Genetics of Nonalcoholic Fatty Liver Disease: Spotlight on PNPLA3 and TM6SF2. Semin Liver Dis 2015;35:270-90. [Crossref] [PubMed]
- 1000 Genomes Project Consortium, Auton A, Brooks LD, et al. A global reference for human genetic variation. Nature 2015;526:68-74.
- Dai G, Liu P, Li X, et al. Association between PNPLA3 rs738409 polymorphism and nonalcoholic fatty liver disease (NAFLD) susceptibility and severity: A meta-analysis. Medicine (Baltimore) 2019;98:e14324. [Crossref] [PubMed]
- Liu YL, Patman GL, Leathart JB, et al. Carriage of the PNPLA3 rs738409 C >G polymorphism confers an increased risk of non-alcoholic fatty liver disease associated hepatocellular carcinoma. J Hepatol 2014;61:75-81. [Crossref] [PubMed]
- Stender S, Kozlitina J, Nordestgaard BG, et al. Adiposity amplifies the genetic risk of fatty liver disease conferred by multiple loci. Nat Genet 2017;49:842-7. [Crossref] [PubMed]
- Grimaudo S, Pipitone RM, Pennisi G, et al. Association Between PNPLA3 rs738409 C>G Variant and Liver-Related Outcomes in Patients with Non-alcoholic Fatty Liver Disease. Clin Gastroenterol Hepatol 2020;18:935-44. [Crossref] [PubMed]
- Luukkonen PK, Nick A, Holtta-Vuori M, et al. Human PNPLA3-I148M variant increases hepatic retention of polyunsaturated fatty acids. JCI Insight 2019. [Crossref] [PubMed]
- Valenti L, Dongiovanni P. Mutant PNPLA3 I148M protein as pharmacological target for liver disease. Hepatology 2017;66:1026-8. [Crossref] [PubMed]
- Linden D, Ahnmark A, Pingitore P, et al. Pnpla3 silencing with antisense oligonucleotides ameliorates nonalcoholic steatohepatitis and fibrosis in Pnpla3 I148M knock-in mice. Mol Metab 2019;22:49-61. [Crossref] [PubMed]
- Shen J, Wong GL, Chan HL, et al. PNPLA3 gene polymorphism and response to lifestyle modification in patients with nonalcoholic fatty liver disease. J Gastroenterol Hepatol 2015;30:139-46. [Crossref] [PubMed]
- Krawczyk M, Stachowska E, Milkiewicz P, et al. Reduction of Caloric Intake Might Override the Prosteatotic Effects of the PNPLA3 p.I148M and TM6SF2 p.E167K Variants in Patients with Fatty Liver: Ultrasound-Based Prospective Study. Digestion 2016;93:139-48. [Crossref] [PubMed]
- Scorletti E, West AL, Bhatia L, et al. Treating liver fat and serum triglyceride levels in NAFLD, effects of PNPLA3 and TM6SF2 genotypes: Results from the WELCOME trial. J Hepatol 2015;63:1476-83. [Crossref] [PubMed]
- Dongiovanni P, Petta S, Mannisto V, et al. Statin use and non-alcoholic steatohepatitis in at risk individuals. J Hepatol 2015;63:705-12. [Crossref] [PubMed]